Group Dr. El-Sherif
Systems Biology of Gene Regulation and Pattern Formation in Development
From Enhancers to Gene Networks
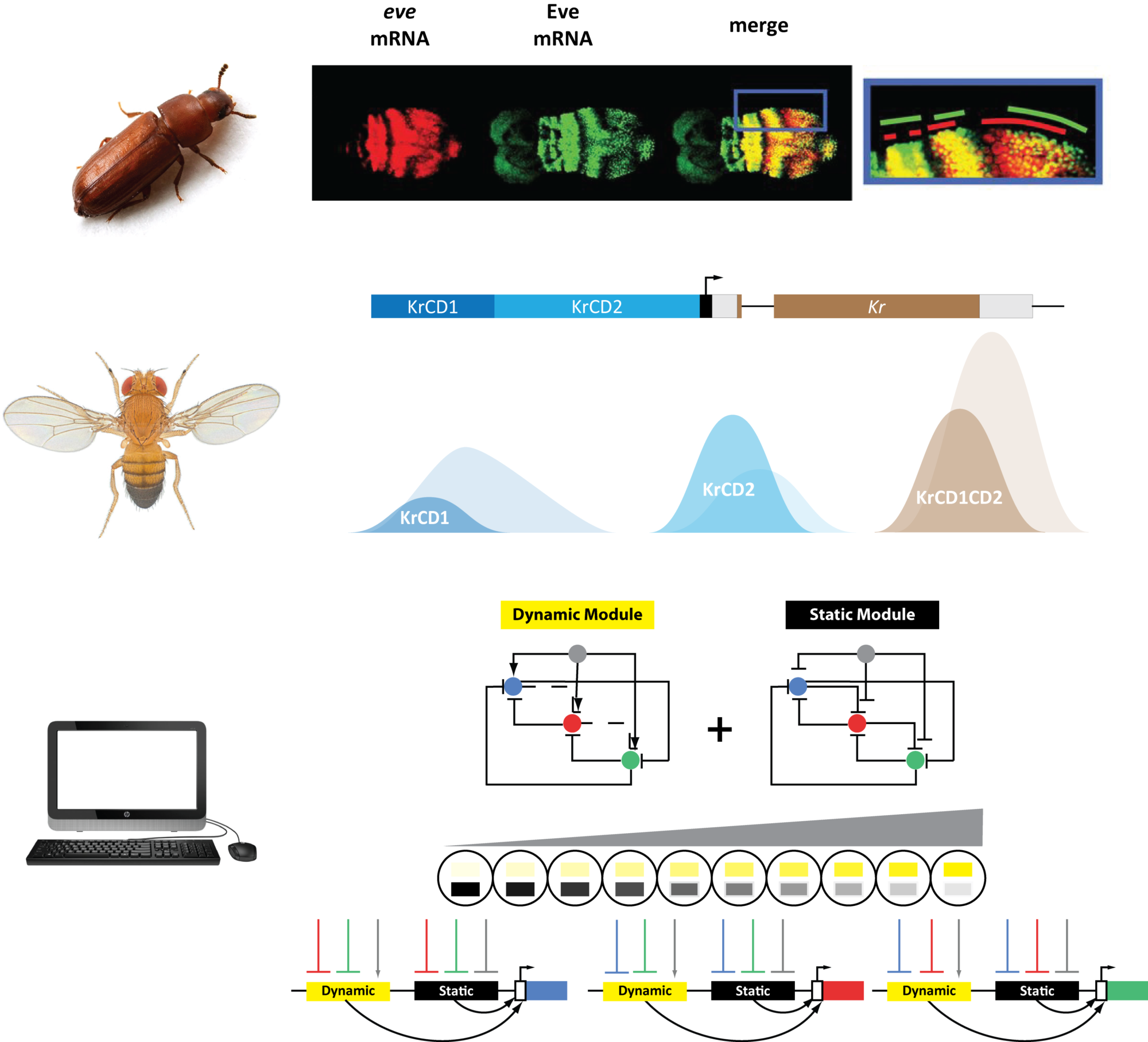
How a homogeneous group of cells is partitioned into domains of different identities is a common problem in embryogenesis. Our group studies how Gene Regulatory Networks (GRNs) act in space and time to generate different patterns (e.g. bands and stripes) of gene expression to partition embryonic tissues into different fates. The main philosophy of our research is the belief that what a GRN can do depends on two factors: (1) the wiring of the GRN (i.e. which gene influences which), and (2) input/output logic of each gene. These two factors are usually blurred because the input/output logic used in GRN modeling is usually a simple and direct realization of the GRN wiring. For example, gene A either activates or represses gene B. However, recent works in cis-regulatory analysis in development found that a gene is usually regulated by many enhancers, each with oftentimes different (and even contradictory) input/output logic. Why using different logics to regulate a single gene and how these different logics are integrated is so far not clear. We believe that the integration of different logics to regulate individual genes within a GRN has an emergent impact on the workings of the GRN as a whole, its robustness, and evolvability. Our group studies this phenomenon both computationally and experimentally. We use phenomenological models (i.e. simple toy models) to try out simple ideas on the computer and generate testable hypotheses. We use the patterning and evolution of anterior-posterior patterning in the beetle Tribolium castaneum and the fruit fly Drosophila melanogaster as model problems to test our ideas experimentally.
Members
Heike Rudolf (PhD student)
Christine Zellner (PhD student)

